|
|
|
|
|
|
| Table 3.
N-Methylethylidenimine. MP2/6-311+G(3d,3p) and B3P86/6-311+G(3d,3p) optimized structure parameters (Å and degrees).
|
|
|
|
|
|
|
Conformer A
|
|
|
|
|
|
|
| 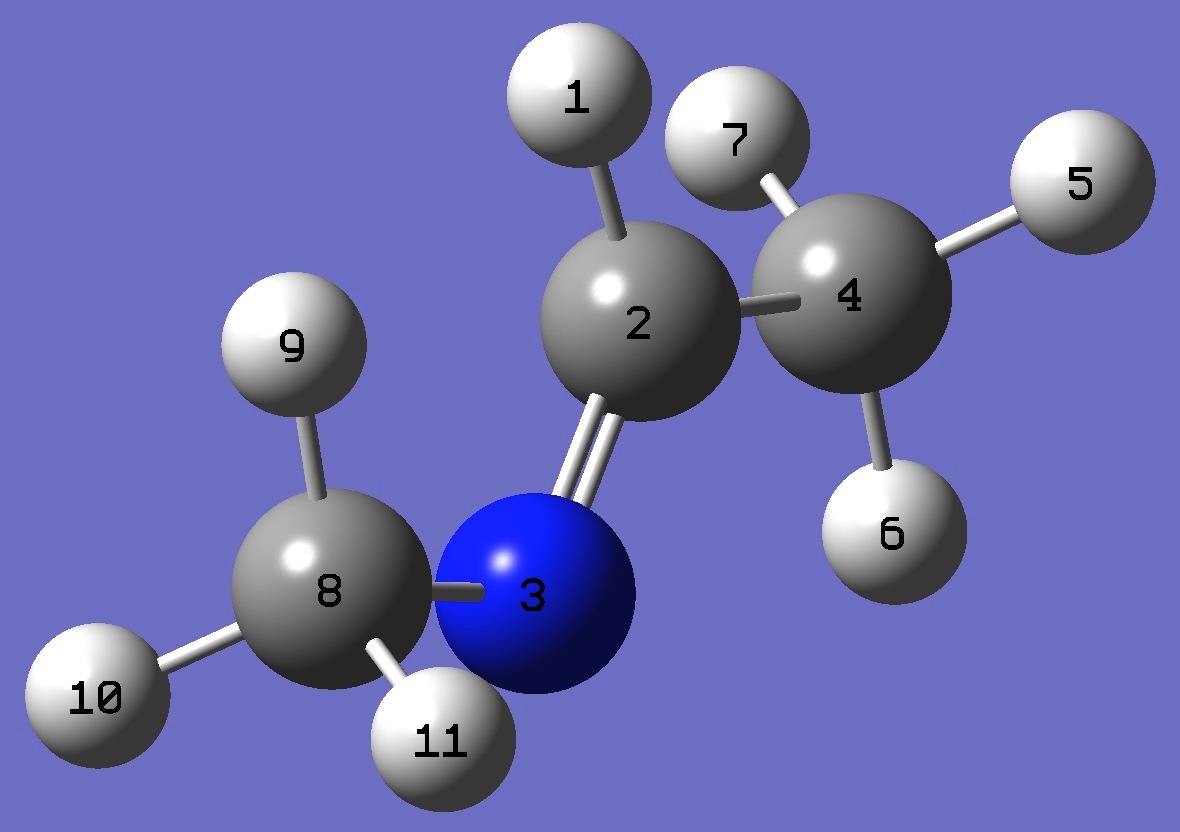 |
H
C,1,B1
N,2,B2,1,A1
C,2,B3,1,A2,3,D1,0
H,4,B4,2,A3,1,D2,0
H,4,B5,2,A4,1,D3,0
H,4,B6,2,A5,1,D4,0
C,3,B7,2,A6,1,D5,0
H,8,B8,3,A7,2,D6,0
H,8,B9,3,A8,2,D7,0
H,8,B10,3,A9,2,D8,0
|
|
|
|
|
|
|
|
|
MP2
|
B3P86
|
|
|
|
|
|
|
|
|
B1=1.09867503
B2=1.27557858
B3=1.49558173
B4=1.09036791
B5=1.08612884
B6=1.09036791
B7=1.4550771
B8=1.0957449
B9=1.08857036
B10=1.08857036
A1=120.95422774
A2=117.09376571
A3=110.27644428
A4=110.09595125
A5=110.27644428
A6=116.81659861
A7=113.13375191
A8=108.93363333
A9=108.93363333
D1=180.
D2=59.08482646
D3=180.
D4=-59.08482646
D5=0.
D6=0.
D7=121.53477697
D8=-121.53477697
|
B1=1.10150248
B2=1.26176601
B3=1.48976364
B4=1.09277791
B5=1.08761052
B6=1.09277791
B7=1.44229573
B8=1.09837409
B9=1.09163771
B10=1.09163771
A1=120.94307496
A2=116.52979244
A3=110.41099341
A4=110.47398196
A5=110.41099341
A6=118.03257793
A7=113.62517162
A8=109.23698563
A9=109.23698563
D1=180.
D2=58.86887781
D3=180.
D4=-58.86887781
D5=0.
D6=0.
D7=121.7916195
D8=-121.7916195
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Conformer B |
|
|
|
|
|
|
|
|
|
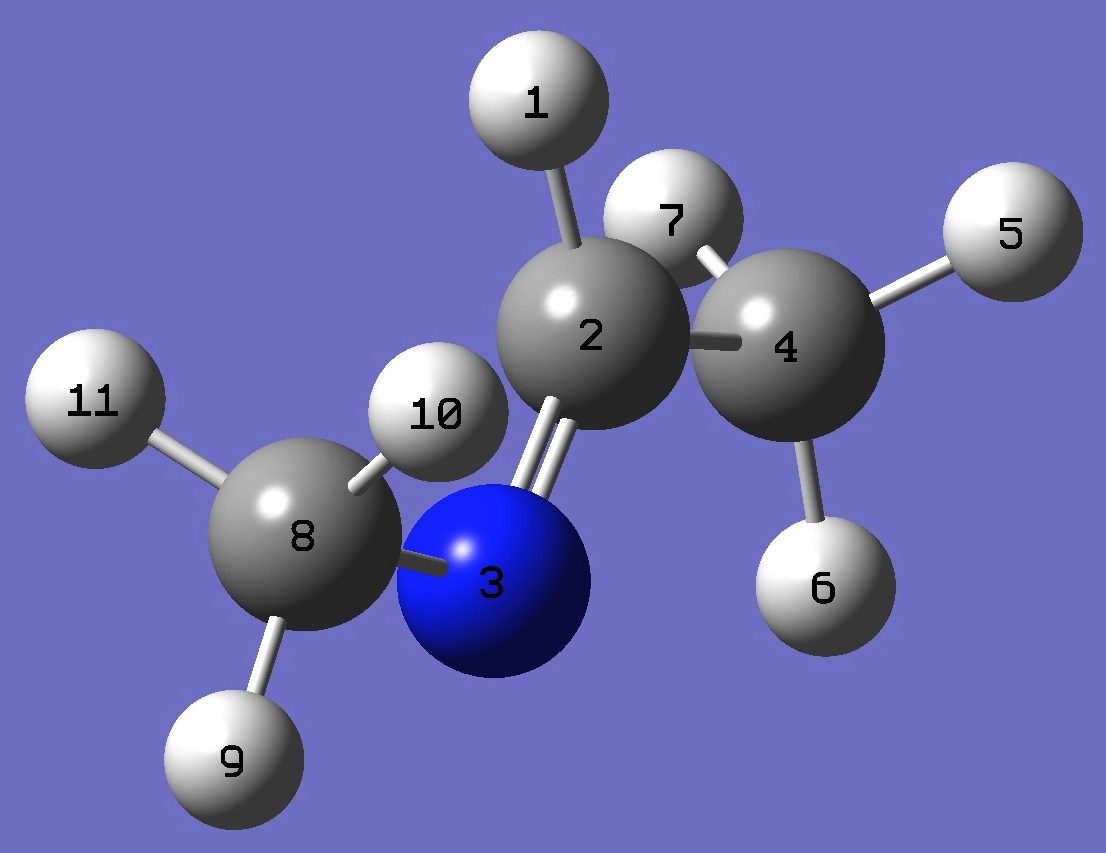
|
H
C,1,B1
N,2,B2,1,A1
C,2,B3,1,A2,3,D1,0
H,4,B4,2,A3,1,D2,0
H,4,B5,2,A4,1,D3,0
H,4,B6,2,A5,1,D4,0
C,3,B7,2,A6,1,D5,0
H,8,B8,3,A7,2,D6,0
H,8,B9,3,A8,2,D7,0
H,8,B10,3,A9,2,D8,0
|
|
|
|
|
|
|
|
MP2
|
B3P86
|
|
|
|
|
|
|
|
|
B1=1.09907705
B2=1.27586595
B3=1.49541325
B4=1.09048763
B5=1.08624325
B6=1.09048763
B7=1.46646725
B8=1.08558134
B9=1.09081467
B10=1.09081467
A1=121.30874769
A2=116.87035206
A3=110.31483284
A4=109.97701774
A5=110.31483284
A6=115.92870607
A7=109.32536233
A8=110.97056506
A9=110.97056506
D1=180.
D2=59.09141852
D3=180.
D4=-59.09141852
D5=0.
D6=180.
D7=-60.43335687
D8=60.43335687
|
B1=1.10166169
B2=1.26184885
B3=1.48976139
B4=1.09293753
B5=1.08769053
B6=1.09293753
B7=1.45380203
B8=1.08829268
B9=1.09350611
B10=1.09350611
A1=121.52781913
A2=116.19566673
A3=110.44116355
A4=110.37203058
A5=110.44116355
A6=117.52333047
A7=109.59993527
A8=111.50658882
A9=111.50658882
D1=180.
D2=58.86407226
D3=180.
D4=-58.86407226
D5=0.
D6=180.
D7=-60.46596605
D8=60.46596605
|
|
|
|
|
|
|
|
|