|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
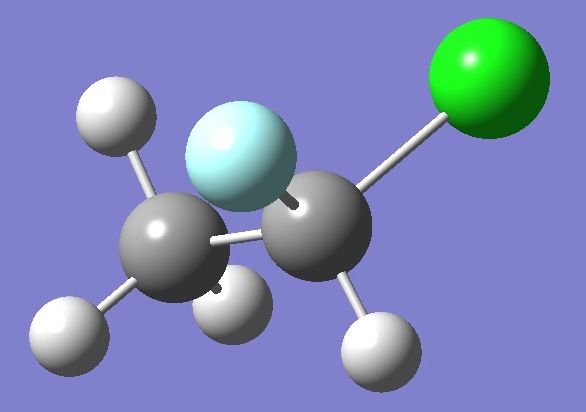
H3C-CHFCl |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Chlorine |
|
|
|
Nuclear
Quadrupole Coupling Constants |
|
|
in 1-Chloro-1-Fluoroethane
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Calculation of the chlorine nqcc's in 1-chloro-1-fluoroethane was made on structures with bond lengths derived ab initio
by the methods of the Lille group, as described below.
Interatomic angles used in the calculation are those given by (1)
MP2/6-311+G(d,p),
and (2) B3P86/6-311+G(3d,3p) optimization. Calculated nqcc's are
compared with the experimental values [1] in Tables 1 and 2.
Structure
parameters are given in Z-Matrix format in Table 3. Atomic
coordinates and rotational constants are given in Tables 4 and 5,
respectively.
|
|
|
|
|
|
|
|
|
|
|
|
|
In Tables 1 and 2, subscripts a,b,c refer to the
principal axes of the inertia tensor; x,y,z to the principal axes
of the nqcc tensor.
Ø (degrees) is the angle between its subscripted
parameters. ETA = (Xxx - Xyy)/Xzz. |
|
|
|
|
|
|
|
|
|
|
|
|
RMS is the root mean square
difference between calculated and experimental diagonal nqcc's
(percentage of the average of the magnitudes of the experimental
nqcc's). RSD is the calibration residual standard deviation of
the B1LYP/TZV(3df,2p) model for calculation of the chlorine nqcc's. |
|
|
|
|
|
|
|
|
|
|
|
|
| |
|
|
|
|
|
|
|
|
|
Table 1. 35Cl
nqcc's in H3C-CHFCl (MHz). Calculation was made on the ab initio structure with interatomic angles given by (1) MP2/6-311+G(d,p), and (2) B3P86/6-311+G(3d,3p) optimization. |
|
| |
|
|
|
|
|
|
|
|
|
|
|
Calc. (1)
|
|
Calc. (2) |
|
Expt. [1] |
|
| |
|
|
|
|
|
|
|
|
|
Xaa |
- |
62.78 |
- |
62.88 |
- |
62.4014(109) |
|
Xbb |
|
34.39 |
|
34.22 |
|
34.2641(138) |
|
Xcc |
|
28.39 |
|
28.66 |
|
28.1373(138) |
|
Xab * |
|
0.61 |
|
0.57 |
|
|
|
|
Xac * |
|
26.69 |
|
26.12 |
|
25.5(64) ** |
|
|
Xbc * |
|
2.01 |
|
1.97 |
|
|
|
|
|
|
|
|
|
|
|
|
|
RMS |
|
0.27 (0.66 %) |
0.41 (0.99 %) |
|
|
|
RSD |
|
0.49 (1.1 %) |
|
0.49 (1.1 %) |
|
|
|
|
|
|
|
|
|
|
|
|
|
Xxx |
|
32.82 |
|
32.74 |
|
|
|
|
Xyy |
|
37.20 |
|
37.07 |
|
|
|
|
Xzz |
- |
70.02 |
- |
69.81 |
|
|
|
|
ETA |
|
0.062 |
|
0.062 |
|
|
|
|
Øz,a |
|
15.17 |
|
14.86 |
|
|
|
|
Øz,b |
|
90.03 |
|
90.03 |
|
|
|
|
Øz,c |
|
105.17 |
|
104.86 |
|
|
|
|
Øz,CCl |
|
0.67 |
|
0.54 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
* The algebraic signs of the
off-diagonal components depend on the orientation of the molecule
with respect to a,b,c axes. Here, the algebraic signs (all
positive) correspond to the atomic a,b,c coordinates given in Table 4. |
|
|
** Absolute value. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| |
|
|
|
|
|
|
|
|
|
Table 2. 37Cl
nqcc's in H3C-CHFCl (MHz). Calculation was made on the ab initio structure with interatomic angles given by (1) MP2/6-311+G(d,p), and (2) B3P86/6-311+G(3d,3p) optimization. |
|
| |
|
|
|
|
|
|
|
|
|
|
|
Calc. (1)
|
|
Calc. (2) |
|
Expt. |
|
| |
|
|
|
|
|
|
|
|
|
Xaa |
- |
49.53 |
- |
49.60 |
- |
49.210(60) |
|
|
Xbb |
|
27.10 |
|
26.97 |
|
26.982(60) |
|
|
Xcc |
|
22.42 |
|
22.64 |
|
22.227(60) |
|
|
Xab * |
|
0.38 |
|
0.33 |
|
|
|
|
Xac * |
|
20.95 |
|
20.50 |
|
|
|
|
Xbc * |
|
1.61 |
|
1.58 |
|
|
|
|
|
|
|
|
|
|
|
|
|
RMS |
|
0.23 (0.70 %) |
0.33 (1.0 %) |
|
|
|
RSD |
|
0.44 (1.1 %) |
|
0.44 (1.1 %) |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
* The algebraic signs of the off-diagonal components depend on the
orientation of the molecule with respect to a,b,c axes. Here, the
algebraic signs (all positive) correspond to the atomic a,b,c
coordinates given in Table 4. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Molecular Structure
|
|
|
|
|
|
|
|
|
|
|
|
|
The molecular structure was optimized
at the MP2/6-311+G(d,p) level of theory.
The optimized CC single bond length was then corrected using the
equation obtained from linear regression analysis of the data given in
Table IX of Ref. [4]. Likewise, the optimized CF bond lengths were
corrected by regression analysis of the data given in Table VI of Ref. [3].
For the CCl bond, the structure was optimized at the MP2/6-311+G(2d,p)
level and corrected by linear regression analysis of the data given in Table
4 of Ref. [2]. The CH bond lengths were corrected using r = 1.001
ropt, where ropt is obtained by MP2/6-31G(d,p) optimization
[5]. Interatomic angles used in the calculation are
those given by (1) MP2/6-311+G(d,p) and (2) B3P86/6-311+G(3d,3p) optimization. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Table 3. H3C-CHFCl Structure parameters (Å
and degrees). Point Group, C1. |
|
|
|
|
|
|
|
|
|
|
Cl |
|
|
|
|
|
|
|
|
C |
1 |
R1 |
|
|
|
|
|
|
C |
2 |
R2 |
1 |
A1 |
|
|
|
|
F |
2 |
R3 |
1 |
A2 |
3 |
D1 |
|
|
H |
2 |
R4 |
1 |
A3 |
3 |
D2 |
|
|
H |
3 |
R5 |
2 |
A4 |
1 |
D3 |
|
|
H |
3 |
R6 |
2 |
A5 |
1 |
D4 |
|
|
H |
3 |
R7 |
2 |
A6 |
1 |
D5 |
|
|
|
|
|
|
|
|
|
|
|
|
MP2 Angles |
B3P86 Angles |
|
|
|
|
|
|
|
|
|
|
|
|
|
R1 |
1.7757 |
|
|
|
|
|
R2 |
1.4992 |
|
|
|
|
|
R3 |
1.3685 |
|
|
|
|
|
R4 |
1.0888 |
|
|
|
|
|
R5 |
1.0879 |
|
|
|
|
|
R6 |
1.0893 |
|
|
|
|
|
|
R7 |
1.0878 |
|
|
|
|
|
|
A1 |
111.22 |
111.44 |
|
|
|
|
A2 |
108.48 |
108.13 |
|
|
|
|
A3 |
106.82 |
105.71 |
|
|
|
|
A4 |
110.10 |
110.13 |
|
|
|
|
A5 |
109.08 |
109.06 |
|
|
|
|
A6 |
109.69 |
110.17 |
|
|
|
|
D1 |
- 120.55 |
- 121.14 |
|
|
|
|
D2 |
124.05 |
123.85 |
|
|
|
|
D3 |
61.17 |
61.04 |
|
|
|
|
D4 |
- 178.83 |
- 179.31 |
|
|
|
|
D5 |
- 58.98 |
- 59.42 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Table 4. H3C-CHF35Cl Atomic coordinates, ropt, MP2 Angles. |
| (More figures are shown than are significant.) |
| |
|
|
|
|
|
|
|
|
|
|
a (Å) |
|
b (Å) |
|
c (Å) |
|
|
|
|
|
|
|
|
|
Cl |
|
1.254019 |
- |
0.034883 |
- |
0.046055 |
|
C |
- |
0.465914 |
- |
0.025623 |
|
0.395365 |
|
C |
- |
1.171873 |
- |
1.241484 |
- |
0.125120 |
|
F |
- |
1.032841 |
|
1.106192 |
- |
0.124619 |
|
H |
- |
0.491449 |
|
0.061864 |
|
1.480344 |
|
H |
- |
0.733212 |
- |
2.140729 |
|
0.302035 |
|
H |
- |
2.222988 |
- |
1.190327 |
|
0.156157 |
|
H |
- |
1.092671 |
- |
1.285933 |
- |
1.209122 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Table 5. H3C-CHF35Cl Rotational constants (MHz). MP2 and B3P86 angles. |
|
|
|
|
|
|
|
MP2 ropt |
B3P86 ropt |
Expt. [1] |
|
|
|
|
|
|
A |
9 112.3 |
9 066.4 |
9 016.268 55(37) |
|
B |
4 687.4 |
4 699.3 |
4 666.701 081(187) |
|
C |
3 352.8 |
3 350.8 |
3 329.369 097(152) |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
[1] R.Hinze, A.Lesarri, J.C.López, J.L.Alonso, and A.Guarnieri, J.Chem.Phys. 104,9729(1996). |
|
|
[2] I.Merke, L.Poteau, G.Wlodarczak,
A.Bouddou, and J.Demaison, J.Mol.Spectrosc. 177,232(1996). |
|
|
[3] R.M.Villamañan, W.D.Chen,
G.Wlodarczak, J.Demaison, A.G.Lesarri, J.C.López, and J.L.Alonso,
J.Mol.Spectrosc. 171,223(1995) |
|
|
[4] J.Demaison, J.Cosléou, R.Bocquet,
and A.G.Lesarri, J.Mol.Spectrosc. 167,400(1994). |
|
|
[5] J.Demaison and G.Wlodarczak, Structural
Chem. 5,57(1994).
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
CH3Cl |
CH3CH2Cl |
CH2ClCHF2 |
CH3CCl3 |
|
|
CF2ClCH3 |
CF2ClCHF2 |
CF2ClCH2F |
CF2ClCF3 |
|
|
CF3Cl |
CH2ClCF3 |
CH2ClCH2F |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Table of Contents |
|
|
|
|
|
Molecules/Chlorine |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
CH3CHFCl.html |
|
|
|
|
|
|
Last
Modified 17 Nov 2005 |
|
|
|
|
|
|
|
|
|
|
